NanoString

NanoString - High Plex, Spatial Profiling of Proteins and RNA from FFPE
Spatial Biology is the study of tissues within their own 2D or 3D context and is the new frontier of molecular biology. In the same way that a GPS captures geographic coordinates within an area to create a map and locates where you are, spatial biology does the same on the tissue, cellular and molecular level.
Through this technology, we can map the spatial architecture of a tissue sample and how cells interact with each other. Additionally, it enables researchers to see multicellular interactions that are not possible by sequencing or any other available technologies.
The NanoString GeoMx® Digital Spatial Profiler (DSP) makes spatial transcriptomics and spatial proteomics possible by visualizing the spatial architecture of intact tissue sections and analyzing multiple biomarkers from a single sample.
GeoMx® Digital Spatial Profiler
Combining standard immunofluorescence techniques with digital optical barcoding technology, the GeoMx® Digital Spatial Profiler enables high plex, spatially resolved profiling of protein and RNA. The GeoMx Assays allow for imaging and profiling from a single FFPE or fresh frozen tissue section.
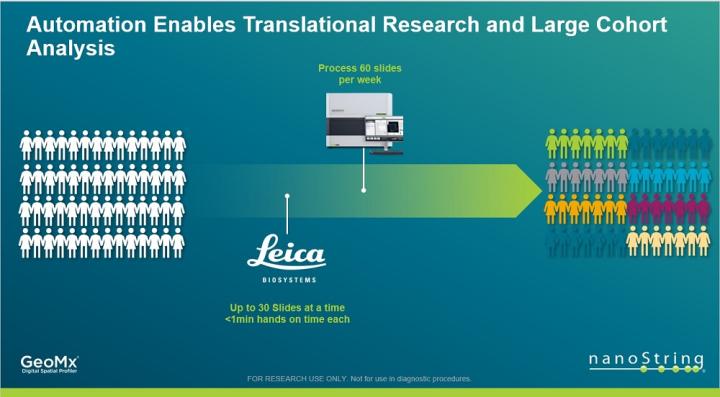
Workflow
Using the BOND RX fully automated research stainer from Leica cuts down on tedious hands-on time staining samples for GeoMx DSP, enabling translational research through batch analysis of large numbers of samples from multi-cohort studies.
Benefits
- Quantify up to 100+ proteins and up to the whole transcriptome with spatial context
- Focus on rare cells or compartment-specific regions of interest in the tissue
- Preserve precious samples with non-destructive processing
- Work with FFPE or fresh frozen tissue
Research Focus
Tissue Atlasing
Deeply profile transcriptomic and proteomic changes in different tissue structures within various organs in order to better understand organ function in health and disease.
Biomarker Discovery and Validation
Discover novel RNA or protein biomarkers that correlate with disease onset, progression, or therapeutic response and are found in specific tissue compartments. Validate these biomarkers in a larger sample set with cohort analysis.
Immune Profiling
Profile the phenotype and transcriptional changes occurring in the immune cell infiltrate within tissues in response to various disease such as cancer, organ rejection, and viral/bacterial infections.
Biology-Driven Profiling: Region of Interest Selection Strategies
Segmentation: Identify and profile distinct biological compartments within a region of interest (ROI).
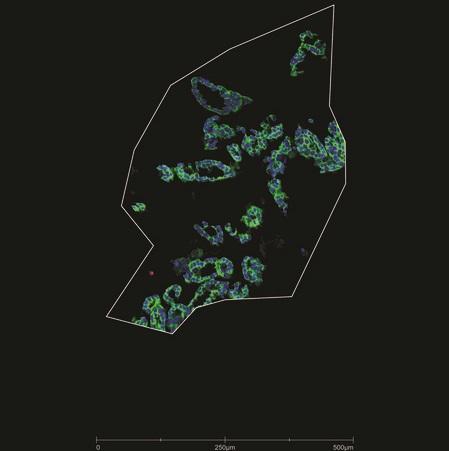
Geometric Profiling: Profile with any geometric shape to characterize distinct tissue regions.
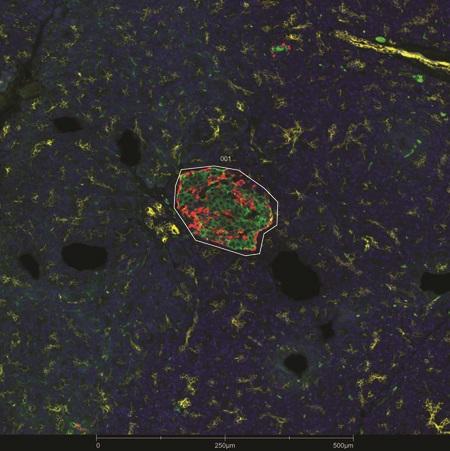
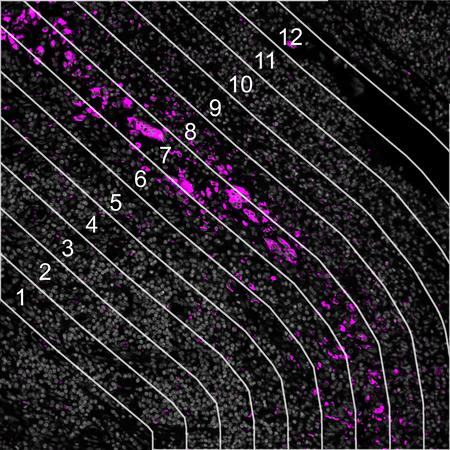
Evaluate how proximity around a central structure affects biological response with radiating ROIs.

Publications
BiTE secretion from in situ-programmed myeloid cells results in tumor-retained pharmacology
| Publication Year: | 2022 |
| Country: | USA |
| Publication: | Hao S, Inamdar VV, Sigmund EC, et al. BiTE secretion from in situ-programmed myeloid cells results in tumor-retained pharmacology. J Control Release. 2022;342:14-25. doi:10.1016/j.jconrel.2021.12.029 |
| Institute: | - |
| Instruments: | Bond Rx, Leica Bond Epitope Retrieval, BOND Polymer Refine Detection DAB, hematoxylin counterstain, nCounter Mouse Myeloid Innate Immunity v2 Panel, nSolver Analysis Software 4.0 |
| Authors: | Hao S, Inamdar VV, Sigmund EC, et al. |
| Study Goals: | To investigate how nanoparticles directs BiTE expression to tumor sites |
| Keywords: | In situ gene therapy, Bi-specific T-cell engagers (BiTEs), Nanotechnology |
System-wide transcriptome damage and tissue identity loss in COVID-19 patients
| Publication Year: | 2022 |
| Country: | USA |
| Publication: | Park J, Foox J, Hether T, et al. System-wide transcriptome damage and tissue identity loss in COVID-19 patients. Cell Rep Med. 2022;3(2):100522. Published 2022 Jan 24. doi:10.1016/j.xcrm.2022.100522 |
| Institute: | Department of Pathology and Laboratory Medicine, Weill Cornell Medicine, New York, NY, USA |
| Instruments: | Leica Biosystems BOND RX, CD45-594 at 1:10 (NanoString Technologies), PanCK-532 at 1:20 (NanoString Technologies) |
| Authors: | Park J, Foox J, Hether T, et al. |
| Study Goals: | To investigate a systemic disruption of canonical cellular and transcriptional pathways across all tissues, which can inform subsequent studies to combat the mortality of COVID-19 and to better understand the molecular dynamics of lethal SARS-CoV-2 and other respiratory infections. |
| Keywords: | Not listed |
For Research Use Only. Not for use in diagnostic procedures.


